Turn on "Hbonds" on the button panel, to see the H-bonds in brown. In KiNG, choose View4 for a close-up from the side, with the helical hydrogen bonds (H-bonds) in brown. Grisham (University of Virginia in Charlottesville, Virginia). This figure is a snaphot of a Java Applet written by Edward K. The amino acid residues are numbered from nearest to most distant and are arranged as an ideal alpha helix with 3.6 residues per complete turn. The figure to the left shows a helical wheel representation of an amino acid sequence, as if looking down the axis of an alpha helix that is perpendicular to the page. In the view menu in KiNG, choose View2 or View3 to see more of the structure. This is a typical globular-protein helix in its native configuration, the polar residues would face the solvent while the hydrophobic residues would face the protein interior. When you clicked the different sidechain types on, what did you observe? Did you notice that the helix has one side with mainly polar residues, and the other with mainly hydrophobic residues?. Now TURN ON and OFF the various display groups and sets, by clicking in the appropriate button box. These can be turned on by clicking on the checkbox labeled "side ch". The hydrophobic side chains are shown in seagreen, polar ones in skyblue, and charged ones in red. Notice that the Cα-Cβ bonds do not point out radially from the helix axis but "pinwheel" along the line of one of the adjacent peptides, giving the side chains an asymmetrical start. The non-integral, 3.6-residue-per-turn repeat of the alpha helix means that the Cα's of successive turns are about halfway offset, giving the main chain a distinctive 7-pointed star appearance in end view. The O and N atoms of the helix main chain are shown as red and blue balls, respectively. The kinemage linked above shows an individual alpha helix, viewed from the N-terminal end to resemble the "helical wheel" (see figure below). If you need information on using King, please hover here.Īn alpha helix is an element of secondary structure in which the amino acid chain is arranged in a spiral. Upon loading this page, the KiNG Java Applet should automatically spawn. To load the KiNG Java Applet, just click here.
#Peptide backbone software
These will be described in the following sections and visualized using the KiNG software mentioned previously. Several common secondary structures have been identified in proteins. While primary structure describes the sequence of amino acids forming a peptide chain, secondary structure refers to the local arrangement of the chain in space. Each amino acid is connected to the next by a peptide bond. By convention, the primary structure of a protein is read and written from the amino-terminal (N) to the carboxyl-terminal (C) end. The primary structure of a peptide or protein is the linear sequence of its amino acids (AAs). Richardson (1992) Protein Science 1: 3-9. Reference: "THE KINEMAGE: A TOOL FOR SCIENTIFIC COMMUNICATION" D.C. Kinemages were originally implemented under the auspices of the Innovative Technology Fund and the Protein Society, and the programming and maintenance are done by David C. These "kinemages" (kinetic images) can be rotated, moved, and zoomed, and parts can be hidden or shown. This module includes links to KiNG (Kinemage, Next Generation), which displays three-dimensional structures in an animated, interactive format. As you traverse though this module please note these aspects. We also view structures in lots of ways - Cα backbone, ball-and-stick, CPK, ribbon, spacefilling - as well color is used to highlight different aspects of the amino acids, structure, etc.
#Peptide backbone series
In this series of pages we examine the different levels of protein organization.
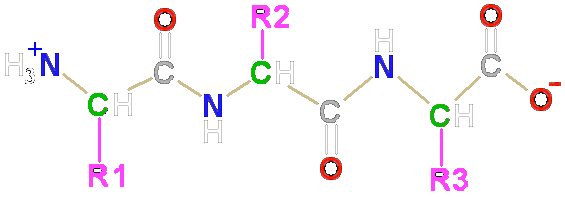
A 2014 Foundations of Medicine eLAB Levels of Protein OrganizationĪ protein's primary structure is defined as the amino acid sequence of its polypeptide chain secondary structure is the local spatial arrangement of a polypeptide's backbone (main chain) atoms tertiary structure refers to the three-dimensional structure of an entire polypeptide chain and quaternary structure is the three-dimensional arrangement of the subunits in a multisubunit protein.
